February 2026
Dating ancient DNA under the multispecies coalescent
“The affinities of all the beings of the same class have sometimes been represented by a great tree. I believe this simile largely speaks the truth. The green and budding twigs may represent existing species; and those produced during each former year may represent the long succession of extinct species.” (Charles Darwin, On the Origin of Species by Means of Natural Selection, Chapter IV, 1859).
The problem of time
Estimating when species diverged is one of the central goals of evolutionary biology. Since molecular sequences accumulate mutations over time, by comparing DNA from different species we can infer how long ago they shared a common ancestor. But there is an inherent limitation: molecular data alone only tell us about relative divergence, not absolute time. This means that without some external reference point, we cannot separate the speed of molecular evolution from the duration over which it has occurred. Researchers have traditionally relied on fossil age calibrations to link phylogenetic trees to an absolute timescale. However, placing fossils on the right branches of a phylogeny is difficult, since they must rely on morphological characters that may be sparse or ambiguous. Ancient DNA (aDNA) offers a different approach. Because aDNA samples come with known ages, from radiocarbon dating or stratigraphic context, they effectively provide timestamps scattered across the branches of a phylogeny. If the time intervals between samples are large enough, or the number of loci is large enough, the expected difference in accumulated mutations between lineages sampled at different times becomes detectable, making it possible to estimate mutation rates and divergence times directly from the data.
When gene trees are not species trees
Our AaRC speaker Dr. Anna Nagel described how she developed a new framework to bring tip dating into the multispecies coalescent (MSC) model, implemented in the widely used program BPP [1]. Why does this matter? Most existing methods that incorporate sample ages into phylogenetic dating, such as BEAST, estimate gene trees rather than species trees. As Anna mentioned, this distinction is critical, since “the common ancestor of a gene must be older than the common ancestor of the species” [1]. As a result, using gene tree divergence times as proxies for species divergence times leads to systematic overestimation. The topologies of different gene trees may also vary, and the coalescence times will almost certainly differ for each locus. The MSC explicitly models gene trees within species trees, accounting for ancestral population sizes, and produces unbiased estimates of species-level divergence times. While the MSC has been available in BPP for years, it previously assumed all sequences were sampled at the present. Anna and her collaborators extended the model to accommodate sequences sampled at different times within each population [1].
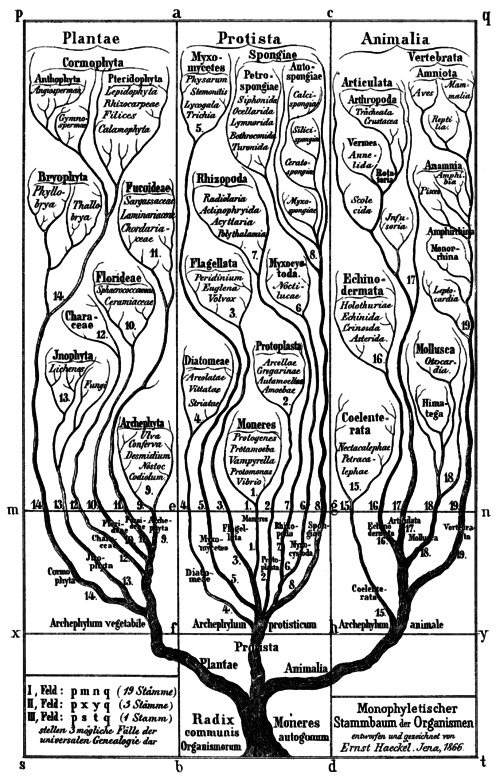
"Monophyletischer Stammbaum der Organismen" (Monophyletic Tree of Organisms). From Ernst Haeckel, Generelle Morphologie der Organismen, Vol. II (Georg Reimer, Berlin, 1866). Via Wikimedia Commons.
What the simulations revealed
Dr. Nagel tested the method extensively through simulations under a range of biologically realistic scenarios. These included populations with divergence times spanning from a few thousand to millions of years. The results were encouraging: with sufficient data, particularly many loci, the method accurately recovered both mutation rates and divergence times. The simulations also showed that the precision of estimates improved most when prioritizing two factors: increasing the number of loci and using older ancient samples. Perhaps unsurprisingly, older samples provide a longer temporal span, making the difference in accumulated mutations more detectable across lineages. One of the most informative parts of the study explored what happens when researchers ignore sample ages and treat all aDNA as contemporary. The consequences were substantial: divergence times were biased, the mutation rate became unidentifiable, and effective population sizes for extinct species were overestimated. These findings serve as a cautionary note for the field, and suggest that even with relatively young aDNA samples, failing to model sample ages properly can lead to misleading inferences.
From mammoths to elephants
To put the method to work on real data, Dr. Nagel analyzed two genomic datasets of elephants and mammoths. One dataset consisted of mitochondrial genomes, including ancient woolly mammoth sequences [2], and the other was a nuclear dataset with sequences from Asian, forest, and savanna elephants, woolly mammoths, and American mastodons [3]. The mitochondrial analysis produced divergence time estimates that were substantially younger than previous estimates, with DNA deamination damage possibly driving inflated mutation rates. This finding highlights a key challenge that the aDNA field still faces: standard damage-filtering approaches may not fully remove the effects of post-mortem modifications, and future work may benefit from explicitly modeling DNA damage itself, as the authors discuss [1]. The nuclear dataset, despite having short loci and limited data, yielded divergence estimates broadly consistent with previous studies that relied on fossil-based calibration. However, the simulations clearly showed that as datasets grow in size, the impact of incorrect sample dates increases, making proper tip dating a relevant issue in the era of large-scale paleogenomic studies.
Looking forward
Dr. Anna Nagel’s work opens several avenues for future development. A practical challenge that remains is the method’s convergence, which the authors found became harder as dataset size increased. Developing better strategies will be important to scale to larger genomic datasets including more loci. The current implementation assumes a fixed species tree topology and no gene flow, assumptions that the latest versions of BPP can relax for contemporary data but not yet for tip-dated analyses. Incorporating migration models would be particularly valuable for systems such as elephants, where hybridization between lineages is well documented, or for hominin datasets where gene flow between archaic and modern populations is a central question. Additionally, combining tip dating with fossil calibrations, and developing models that explicitly account for DNA degradation, could further improve the accuracy and utility of this framework.
References
- Nagel, A.A. et al. Bayesian inference under the multispecies coalescent with ancient DNA sequences. Syst Biol 73: 964–978 (2024).
- van der Valk, T. et al. Million-year-old DNA sheds light on the genomic history of mammoths. Nature 591: 265–269 (2021).
- Rohland, N. et al. Genomic DNA sequences from mastodon and woolly mammoth reveal deep speciation of forest and savanna elephants. PLoS Biol 8: e1000564 (2010).
Below, Anna shared with us further details about her profile, career, prospects and future projects:
1. Briefly introduce yourself. What is your origin story for how you got into science?
I am a postdoc Washington University in St. Louis. I work on population genetics and phylogenetics methods development, mostly relating to the estimation of times. I was always excited about science, but I started as a chemistry major. I was interested in Ecology from spending time at an environmental learning center as a kid, but I thought biology wasn’t mathematical enough. After taking Evolution, I realized that wasn’t true and mathematical models are a powerful tool for applications ranging from proof of concept to inference. Moreover, I was really excited by how studying evolutionary theory allows us to understand the patterns of biological diversity across scales, from individual loci to the tree of life.
2. How and/or why did you start working on this project?
I was actually thinking about HIV transmission and how coalescent models could be used to model transmission with multiple sequences from each infected individual. That project would have required a lot of additions to existing software, including adding tip dating to the multispecies coalescent (MSC). I decided to work on aDNA first because it was a much smaller project. I still haven’t gotten back to the HIV transmission project.
3. Were there any major challenges in this project? How did you overcome them?
One of the main obstacles was getting the MCMCs to mix. Addressing this required developing MCMC proposals specific to tip dating.
4. What do you think are the main take-home messages of this project?
Even relatively young age sequences (in comparison to the species divergence times) can give a lot of information about species divergence times and allow us to estimate those times without outside information such as fossil ages. However, DNA degradation may be a pervasive problem in these types of analyses and future work should explicitly model DNA degradation.
5. What do you think is missing in the field that you would like to work on?
I would be excited to work on migration models with aDNA. Bayesian inference with coalescent models with migrations are difficult to get to work correctly because they are really complex. However, these models would allow us to analyze a more datasets more satisfactorily. In my empirical analysis, I removed a mammoth species that had been suggested to have been a product of hybrid speciation, and likely my estimate of divergence time estimate of the African elephant species was biased substantially due to hybridization. Hominid datasets would also require migration models.
6. Where do you see yourself in the near future?
Right now, I’m hoping to keep working on similar types of questions relating to Bayesian phylogenetic inference and the intersection of phylogenetics and population genetics. I’ve also been working on machine learning in phylogenetics, so I’d like to see how those methods could be applied to the MSC.

